Learn you Rake
Rake is a task runner, and in my opinion, a worthy replacement of Make. It is written in Ruby and, as a result, has the immense benefit of being concise, pleasant to eyes, and fun to write.
Read more →Vim for Writing
I happen to be a Vim user—not because I practice terminal voodoo or intend to push the limits of keyboard efficiency—because Vim is what I started with and had no reasons to move on. If you do not find Vim to be a reasonable choice, that is perfectly fine. There exists a natural equilibrium in the community where users keep keep updating their preferences as the tools evolve.
Read more →Open {data, science, source} - OpenSNP.org
April, 2016
“What’s the worst that can happen? Not like it will be different this time. some sleep”, I told myself as I prepared to brace the bad news. The clock moves closer to midnight with every passing moment, but I am too tired to think about it anymore.
Read more →Quick Data Plotting in R - II
In the first part of the posts on R, I outlined a couple of useful strategies for quick exploratory data analysis and visualization using R programming language. In this post, I will share a few more examples (in a case-study like manner) along with relevant plots for better demonstration.
Read more →Quick Data Plotting in R - I
Python is my all time favorite programming language. I use it all the time. It is simple, readable and easy to get started, something which I picked up four years ago when I went through the first offering of Interactive Programming with Python course on Coursera. Python’s simplicity and appeal has continued to grow over time, supported by a wide community and a rich environment of packages.
Read more →WIGI – an inspire grantee
The Wikipedia Indicators of Gender Inequality (WIGI) project aims to study and characterize gender representation on Wikipedia by studying trends in growth of biography articles. The project is led by Maximilian Klein, a PhD student at the University of Minnesota, and Dr. Piotr Konieczny.
Read more →Bypassing Proxy Servers
IIT Kharagpur uses a proxy server to enable outside internet connections. Whatever be the use case from the perspective of the institution, it is a nightmare to use for us students. Most of the standard ports are blocked resulting in SSH difficulties, accessing ill-fated websites, or inability to use programs that connect directory to a specific address or port. Further, not all applications honor the proxy settings further increasing your frustration, specially if you are a programmer.
Read more →Diving Deeper into BLAST - III
BLAST+ is an incredible tool and does it job in a pretty decent way. However, anyone who has worked with it for some time with some non trivial details has their own horror stories to tell. For the past few months where I have been engaged in development of SequenceServer, we got a lot of opportunities to see the ugly inconsistencies buried within an otherwise beautiful program.
Read more →Diving Deeper into BLAST - II
This is a follow up of the
previous post
where I was trying to parse the BLAST+ XML output to create an efficient data
layer in SequenceServer. A critical and often demanded feature for the
application was the ability to have a graphical overview of all the obtained
hits. NCBI’s BLAST portal, for instance, includes a graphical overview of
your results that summarizes how many hits you got and how do they score.
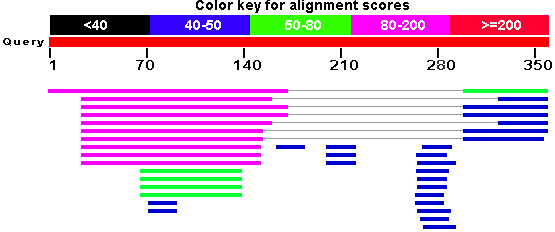 Graphical summary of BLAST results as displayed on NCBI portal.
However, we had nothing for SequenceServer.
Graphical summary of BLAST results as displayed on NCBI portal.
However, we had nothing for SequenceServer.
Diving Deeper into BLAST - I
In my previous post, I mentioned about a project I was trying to work on: SequenceServer. Also, in the end I said that I would be writing about the BLAST algorithm (which is the backbone of this project) and how does it works so efficiently in producing alignments even with very long sequences. However, In this post I would like to talk about BLAST program and it's output before we go into the algorithm some time later.
Read more →Introduction to Sequenceserver
Contributing to open source is quite an exciting journey. I embarked upon this path recently with few contributions to SequenceServer - A BLAST tool, a bioinformatics software built around BLAST+ that lets you setup a custom BLAST+ server to perform BLAST queries with your own database locally through a web interface.
Read more →